Translational mitochondrial pharmacology
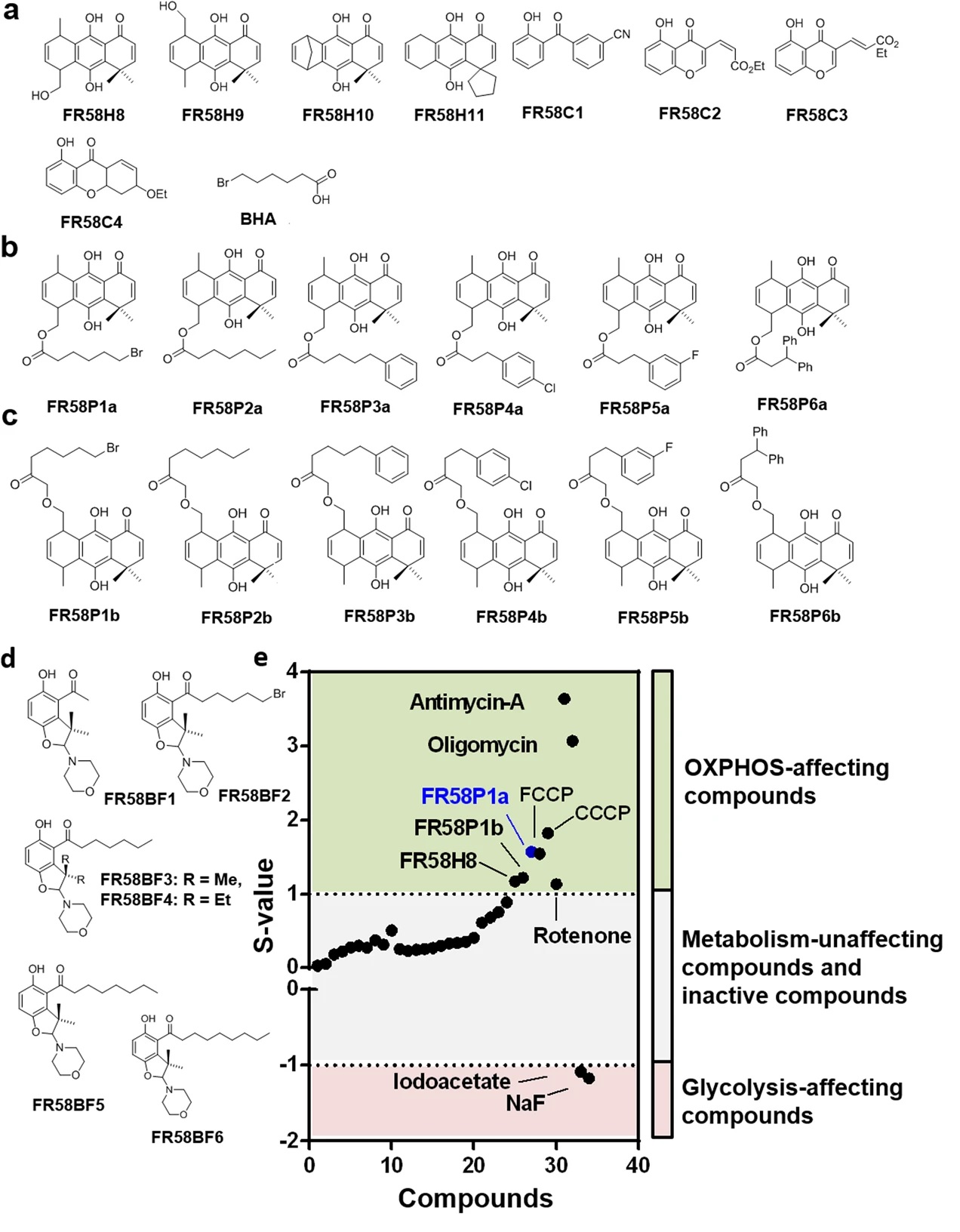
We investigate mitochondrial function as a pharmacological target and as a source of therapeutic innovation. This line focuses on identifying and characterizing small molecules, bioactive compounds, and mitochondria-directed strategies that interfere with key bioenergetic processes relevant to disease.
Our goal is to discover and evaluate compounds with potential anti-cancer, anti-migratory, or anti-platelet effects, while also understanding how mitochondrial metabolism shapes drug response and cellular vulnerability.
Cancer metabolic plasticity
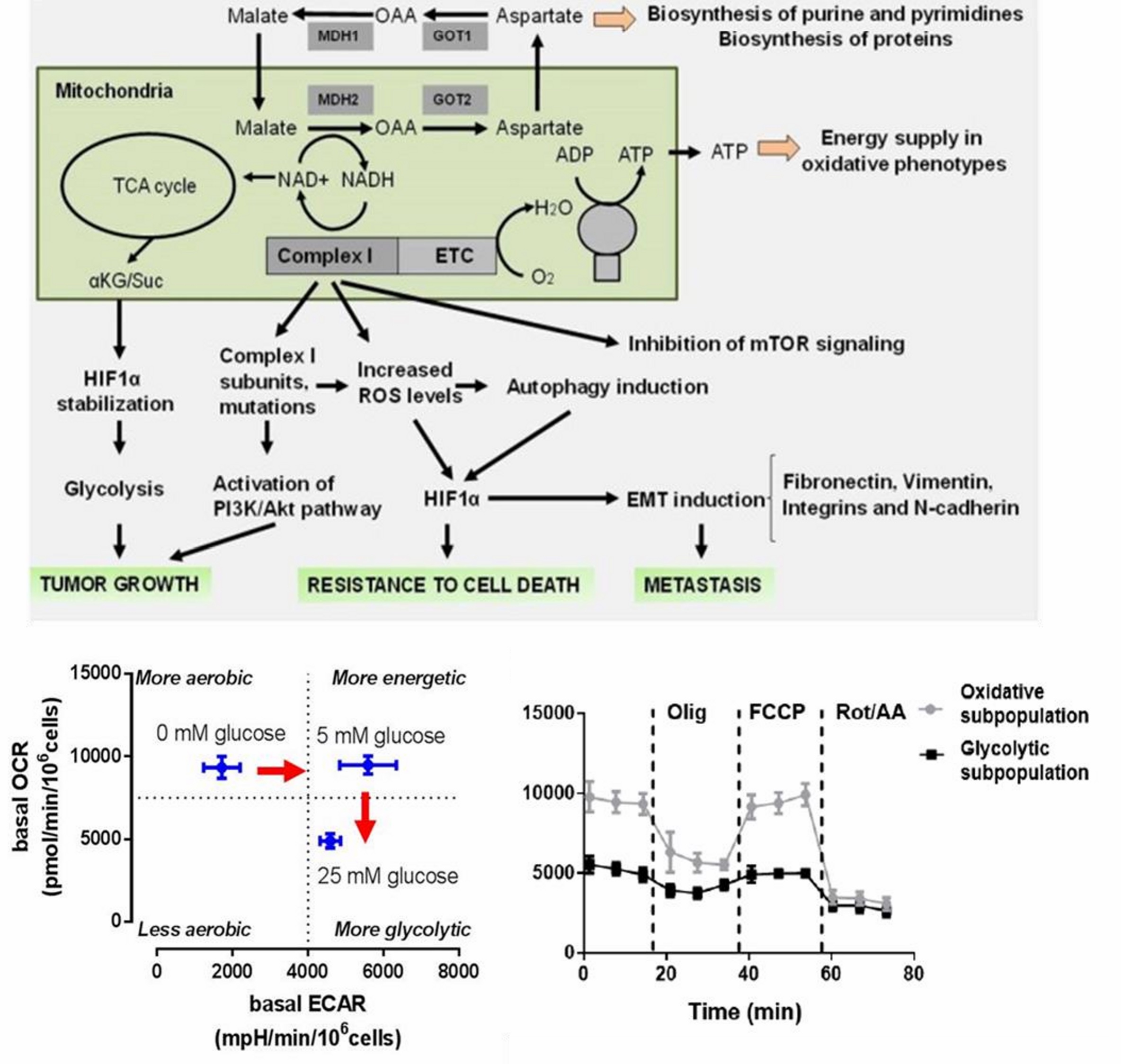
Cancer cells continuously adapt their metabolism to support proliferation, migration, invasion, and survival under therapeutic stress. We study how mitochondrial bioenergetics is rewired during these transitions, with particular interest in the roles of calcium signaling, substrate use, redox balance, and metabolic flexibility.
By studying these adaptive responses in real time, we aim to identify metabolic vulnerabilities that could support new strategies to counter tumor progression, chemoresistance, and metastasis.
Oncovenomics
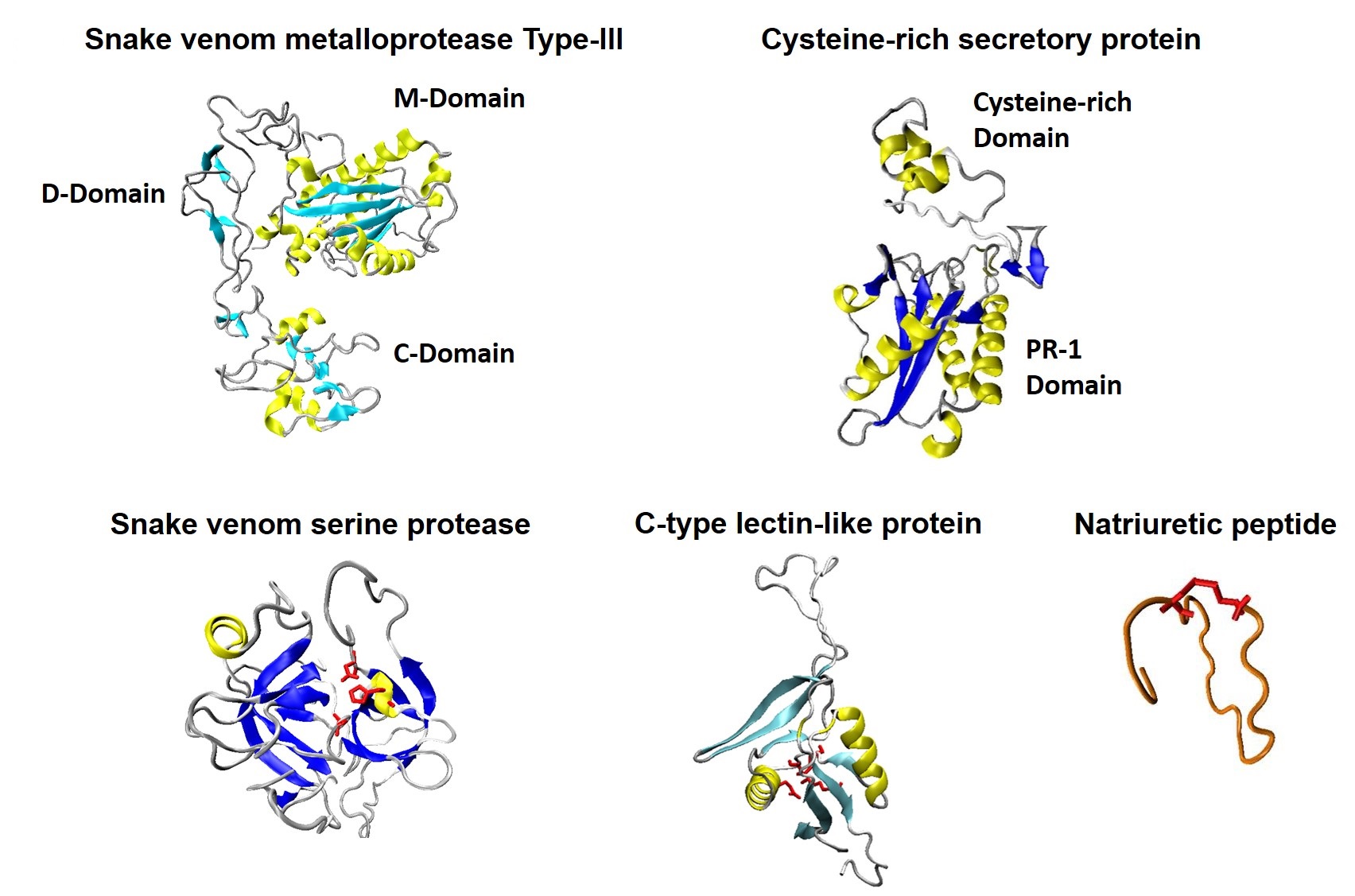
Venoms are rich sources of biologically active molecules with unexplored therapeutic potential. In this line, we study toxins and toxin-inspired molecules from snake venoms and other natural secretions as tools to understand mitochondrial regulation and as candidates for drug discovery in oncology.
Our work in oncovenomics combines molecular, cellular, and functional approaches to characterize mechanisms of action, identify novel bioactivities, and explore how venom-derived compounds may interfere with cancer cell metabolism, migration, and survival.
Museomics and applied herpetology

Our research also integrates biodiversity, herpetology, and molecular approaches to study medically relevant snakes and their toxins. This includes work on venom systems, toxin-encoding genes, comparative biology, and the biomedical and ecological implications of snake diversity in Chile and South America.
Through biological collections, evolutionary questions, and translational toxinology, we create a bridge between biodiversity-based science and biomedical discovery.
